The usual way of running REANA workflows is by means of the command-line interface (CLI) client. In this blog post, we present a step-by-step guide on how to use the Python API instead. This way of interacting with REANA platform can be useful if you would like to integrate REANA workflow submissions with your Python application or simply use your programmable Python environment instead of the command line.
In the following example, we shall use the ROOT6 Roofit example analysis. We shall create a new workflow, upload analysis code and inputs, start the workflow, observe its progress status, retrieve workflow logs, list workspace files and finally download and plot results.
Installing prerequisites
The main prerequisite is to install reana-client. We shall use the REANA 0.7 version in this example:
$ pip install --user reana-client==0.7.2
We shall also need Python imaging library Pillow to display analysis
output plots:
$ pip install --user Pillow
Configuring access
As a first step, we shall configure the environment variable
REANA_SERVER_URL. This variable is mandatory and should point to
the remote REANA cluster instance that will be used to run our
computations. The client code will use this environment variable
automatically to connect to the remote REANA cluster later on. The
typical value is https://reana.cern.ch for CERN deployments. Here is
how we can set it:
import os
if not os.getenv('REANA_SERVER_URL'):
os.environ['REANA_SERVER_URL'] = 'https://reana.cern.ch'
We also need another environment variable, REANA_ACCESS_TOKEN, that
contains our personal user access token. One way to obtain the access
token is by opening https://reana.cern.ch in your browser, signing in
with your CERN credentials and navigating to your user profile. Please
remember to keep your access token private and safe, and please do not
share it with anybody.
from getpass import getpass
my_reana_token = \
os.getenv("REANA_ACCESS_TOKEN") or getpass('Enter your REANA token: ')
Verifying access
Let us verify whether we can well connect to the REANA cluster. In
command-line interface, we would use the
ping
CLI command. In Python, the corresponding function to use is ping():
from reana_client.api.client import ping
ping(my_reana_token)
Example output:
{
'email': 'john.doe@example.org',
'full_name': 'John Doe',
'reana_server_version': '0.7.2',
'reana_token': {
'requested_at': 'Mon, 01 Mar 2021 9:00:00 GMT',
'status': 'active',
'value': 'my_reana_token'
},
'username': None,
'status': 'Connected',
'error': False
}
Specifying workflow
Now that we can connect to the remote REANA cluster, let us proceed to creating our workflow. As a first step, we shall choose a suitable name for our workflow:
my_workflow_name = 'root6-roofit'
We proceed to declaring our workflow inputs. The analysis inputs
consist of two C++ code files and some input parameters, such as the
number of events to generate and the desired output file names. Please
refer to
reana.yaml
for correspondence. We define my_inputs in the following way:
my_inputs = {
'files': [
'code/gendata.C',
'code/fitdata.C'
], # A list of files your analysis will be using
'parameters': {
'events': '20000',
'data': 'results/data.root',
'plot': 'results/plot.png',
} # Parameters for your workflow
}
We now have to specify the computational steps that are necessary to
perform the analysis and obtain our results. The analysis consists of
two steps, the gendata step, where we generate the signal and
background data, and the fitdata step, where we fit the data against
a model. Each step runs in a containerised ROOT environment. Here is
how we can define the workflow steps. Again, please refer to
reana.yaml
for correspondence:
my_workflow = {
'steps': [
{
'name': 'gendata',
'environment': 'reanahub/reana-env-root6:6.18.04',
'commands': [
'mkdir -p results',
'root -b -q \'code/gendata.C(${events},"${data}")\' | tee gendata.log',
],
},
{
'name': 'fitdata',
'environment': 'reanahub/reana-env-root6:6.18.04',
'commands': [
'root -b -q \'code/fitdata.C("${data}","${plot}")\' | tee fitdata.log'
],
},
]
}
Our workflow is linear, so we shall use the simple serial workflow engine that will execute the given steps sequentially:
my_workflow_type = 'serial'
Creating workflow
We are now ready to create our fully-specified workflow in the REANA platform. In command-line interface, we would use the create CLI command. In Python, the corresponding function to use is create_workflow_from_json() function, where we shall pass our previously-created parameters:
from reana_client.api.client import create_workflow_from_json
create_workflow_from_json(
workflow_json=my_workflow,
name=my_workflow_name,
access_token=my_reana_token,
parameters=my_inputs,
workflow_engine=my_workflow_type)
Example output:
{
'message': 'Workflow workspace created',
'workflow_id': '6cd613eb-f2fb-411b-9601-c89599925759',
'workflow_name': 'root6-roofit'
}
Uploading files
We can now proceed to uploading our input files to the workflow workspace. In command-line interface, we would use the upload CLI command. In Python, the corresponding function to use is upload_to_server() that we can call as follows:
from reana_client.api.client import upload_to_server
abs_path_to_input_files = [os.path.abspath(f) for f in my_inputs['files']]
upload_to_server(my_workflow_name, abs_path_to_input_files, my_reana_token)
Starting workflow
At this point we are able to start our workflow in REANA. In command-line interface, we would use the start CLI command. In Python, the corresponding function to use is start_workflow() function.
We shall pass our workflow name and access token. Additionally, we can override workflow parameters while starting the workflow, for example to specify a different number of events that we would like to generate. For now, let us keep the original input parameters, so that we shall pass an empty dictionary:
from reana_client.api.client import start_workflow
start_workflow(my_workflow_name, my_reana_token, {})
Example output:
{
'message': 'Workflow submitted.',
'run_number': 1,
'status': 'queued',
'user': '1d27737b-5b80-4da2-87bb-816a20e110bd',
'workflow_id': '6cd613eb-f2fb-411b-9601-c89599925759',
'workflow_name': 'root6-roofit'
}
Checking workflow status
We are now interested to check the workflow status. Is it waiting in the queue? Is it running? Has it failed or finished? In command-line interface, we would use the status CLI command. In Python, the corresponding function to use is get_workflow_status() function.
Since our example workflow may take several minutes to finish all the computations, we shall use a loop to print its status regularly after a certain wait time:
import time
from reana_client.api.client import get_workflow_status
while True:
status_details = get_workflow_status(my_workflow_name, my_reana_token)
print('Current status: ', status_details['status'])
if status_details['status'] == 'finished':
break
time.sleep(10)
Example output:
Current status: running
...
Current status: finished
Checking workflow logs
After our workflow finishes, or anytime when we would like to debug and investigate how the workflow run, we may want to display the logs of the concrete computational steps. In command-line interface, we could use the logs CLI command. In Python, the corresponding function to use is get_workflow_logs() function.
We need to specify only the workflow name and the access token. Optionally, we can add parameters such as list of steps the logs of which we are interested to see, or pagination parameters in case the log output is too large or too verbose. As an example, let us retrieve the last 15 lines of the “fitdata” step:
import json
from reana_client.api.client import get_workflow_logs
workflow_logs = get_workflow_logs(my_workflow_name, my_reana_token, ['fitdata'])
job_logs = json.loads(workflow_logs['logs'])['job_logs']
fitdata_logs = job_logs[next(iter(job_logs))]['logs']
for item in fitdata_logs.split('\n')[-15:]:
print(item)
Example output:
ERR MATRIX APPROXIMATE
PARAMETER CORRELATION COEFFICIENTS
NO. GLOBAL 1 2 3 4 5
1 0.00000 1.000 0.000 0.000 0.000 0.000
2 0.00000 0.000 1.000 0.000 0.000 0.000
3 0.00000 0.000 0.000 1.000 0.000 0.000
4 0.00000 0.000 0.000 0.000 1.000 0.000
5 0.00000 0.000 0.000 0.000 0.000 1.000
ERR MATRIX APPROXIMATE
[#1] INFO:Minization -- RooMinimizer::optimizeConst: deactivating const optimization
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) directly selected PDF components: (bkg)
[#1] INFO:Plotting -- RooAbsPdf::plotOn(model) indirectly selected PDF components: ()
Info in <TCanvas::Print>: png file results/plot.png has been created
Listing workspace files
We are usually interested in listing the output files which our workflow produced in its workspace. In command-line interface, we would use the ls CLI command. In Python, the corresponding function is list_files() that operates also on the workflow name:
from reana_client.api.client import list_files
list_files(my_workflow_name, my_reana_token)
Example output:
[
{
'last-modified': '2021-03-01T08:57:01',
'name': 'code/gendata.C',
'size': 1937,
},
{
'last-modified': '2021-03-01T08:57:01',
'name': 'code/fitdata.C',
'size': 1648,
},
{
'last-modified': '2021-03-01T08:57:52',
'name': 'results/plot.png',
'size': 15450,
},
{
'last-modified': '2021-03-01T08:57:42',
'name': 'results/data.root',
'size': 154458,
},
]
Downloading outputs
We are now interested in downloading workflow outputs, or any temporary files from the workflow’s workspace, for closer inspection. In command-line interface, we would use the download CLI command. In Python, the corresponding function is download_file() function.
Let us download a binary blob of the final plot which our workflow produced:
from reana_client.api.client import download_file
output_filename = 'results/plot.png'
file_binary_blob = download_file(
my_workflow_name, output_filename, my_reana_token)
Displaying output images
We can now display the final fit produced by our workflow:
from PIL import Image
import io
image_stream = io.BytesIO(file_binary_blob)
img = Image.open(image_stream)
img.show()
Example output:
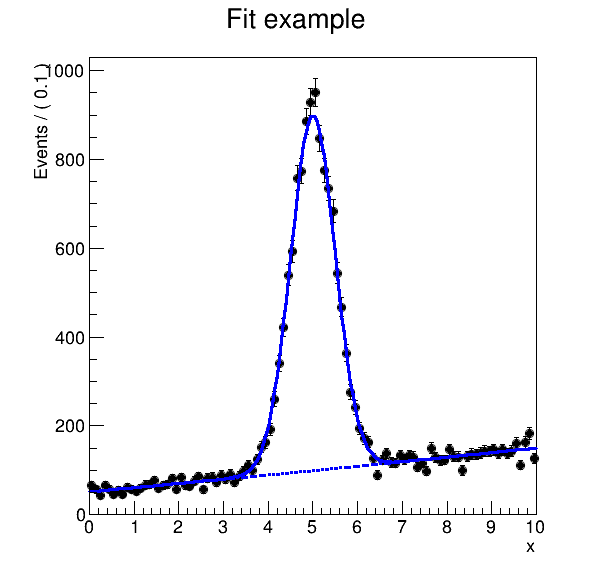
See also: